Note
Go to the end to download the full example code
Differentially Private SGD.
Differentially Private Stochastic Gradient Descent (DP-SGD) [Abadi et al., 2016] is a rather simple idea. Instead of doing a typical SGD update, with the sum of all gradients in a minibatch,
DP-SGD first truncates each individual gradient if their norm exceeds some threshold \(C\) to ensure no single example influence the overall update too much,
and adds Gaussian noise to the update,
That’s the TL:DR, anyway.
It is not too difficult to get to an implementation of DP-SGD that works,
but getting individual gradients from a minibatch in a way that scales can be
tricky.
This examples shows how to use the BatchGrad extension,
which gives access to individual gradients.
Let’s get the imports, configuration and some helper functions out of the way first.
import matplotlib.pyplot as plt
import torch
import torch.nn as nn
from torch.optim import Optimizer
from backpack import backpack, extend
from backpack.extensions import BatchGrad, BatchL2Grad
from backpack.utils.examples import get_mnist_dataloader
NUM_EPOCHS = 1
PRINT_EVERY = 50
MAX_ITER = 200
BATCH_SIZE = 64
DEVICE = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
torch.manual_seed(0)
def make_broadcastable(v, X):
"""Returns a view of `v` that can be broadcast with `X`.
If `v` is a one-dimensional tensor [N] and `X` is a tensor of shape
`[N, ..., ]`, returns a view of v with singleton dimensions appended.
Example:
`v` is a tensor of shape `[10]` and `X` is a tensor of shape `[10, 3, 3]`.
We want to multiply each `[3, 3]` element of `X` by the corresponding
element of `v` to get a matrix `Y` of shape `[10, 3, 3]` such that
`Y[i, a, b] = v[i] * X[i, a, b]`.
`w = make_broadcastable(v, X)` gives a `w` of shape `[10, 1, 1]`,
and we can now broadcast `Y = w * X`.
"""
broadcasting_shape = (-1, *[1 for _ in X.shape[1:]])
return v.reshape(broadcasting_shape)
def accuracy(output, targets):
predictions = output.argmax(dim=1, keepdim=True).view_as(targets)
return predictions.eq(targets).float().mean().item()
Creating the model and loading some data
We will use a small CNN with 2 convolutions, 2 linear layers, and feed it some MNIST data.
def make_small_cnn(outputs=10, channels=(16, 32), fc_dim=32, kernels=(8, 4)):
return nn.Sequential(
nn.ZeroPad2d((3, 4, 3, 4)),
nn.Conv2d(1, channels[0], kernels[0], stride=2, padding=0),
nn.ReLU(),
nn.MaxPool2d(2, stride=1),
nn.Conv2d(channels[0], channels[1], kernels[1], stride=2, padding=0),
nn.ReLU(),
nn.MaxPool2d(2, stride=1),
nn.Flatten(),
nn.Linear(channels[1] * 4 * 4, fc_dim),
nn.ReLU(),
nn.Linear(fc_dim, outputs),
)
mnist_dataloader = get_mnist_dataloader(batch_size=BATCH_SIZE)
model = make_small_cnn().to(DEVICE)
loss_function = nn.CrossEntropyLoss().to(DEVICE)
and we need to extend the model so that BackPACK knows about it.
Computing clipped individual gradients
Before writing the optimizer class, let’s see how we can use BackPACK
on a single batch to compute the clipped gradients, without the overhead
of the optimizer class.
We take a single batch from the data loader, compute the loss,
and use the with(backpack(...)) syntax to activate two extensions;
BatchGrad and BatchL2Grad.
x, y = next(iter(mnist_dataloader))
x, y = x.to(DEVICE), y.to(DEVICE)
loss = loss_function(model(x), y)
with backpack(BatchL2Grad(), BatchGrad()):
loss.backward()
BatchGrad computes individual gradients and BatchL2Grad their norm (squared),
which get stored in the grad_batch and batch_l2 attributes of the parameters
for p in model.parameters():
print(
"{:28} {:32} {}".format(
str(p.grad.shape), str(p.grad_batch.shape), str(p.batch_l2.shape)
)
)
torch.Size([16, 1, 8, 8]) torch.Size([64, 16, 1, 8, 8]) torch.Size([64])
torch.Size([16]) torch.Size([64, 16]) torch.Size([64])
torch.Size([32, 16, 4, 4]) torch.Size([64, 32, 16, 4, 4]) torch.Size([64])
torch.Size([32]) torch.Size([64, 32]) torch.Size([64])
torch.Size([32, 512]) torch.Size([64, 32, 512]) torch.Size([64])
torch.Size([32]) torch.Size([64, 32]) torch.Size([64])
torch.Size([10, 32]) torch.Size([64, 10, 32]) torch.Size([64])
torch.Size([10]) torch.Size([64, 10]) torch.Size([64])
To compute the clipped gradients, we need to know the norms of the complete individual gradients, but ad the moment they are split across parameters, so let’s reduce over the parameters
l2_norms_squared_all_params = torch.stack([p.batch_l2 for p in model.parameters()])
l2_norms = torch.sqrt(torch.sum(l2_norms_squared_all_params, dim=0))
We can compute the clipping scaling factor for each gradient,
given a maximum norm C,
as a tensor of [N] elements.
C = 0.1
scaling_factors = torch.clamp_max(l2_norms / C, 1.0)
All that remains is to multiply the individual gradients by those factors and sum them to get the update direction for that parameter.
for p in model.parameters():
clipped_grads = p.grad_batch * make_broadcastable(scaling_factors, p.grad_batch)
clipped_grad = torch.sum(clipped_grads, dim=0)
Writing the optimizer
Let’s do the same, but in an optimizer class.
class DP_SGD(Optimizer):
"""Differentially Private SGD.
Arguments:
params (iterable): iterable of parameters to optimize or dicts defining
parameter groups
lr (float, optional): coefficient that scale delta before it is applied
to the parameters (default: 1.0)
max_norm (float, optional): maximum norm of the individual gradient,
to which they will be clipped if exceeded (default: 0.01)
stddev (float, optional): standard deviation of the added noise
(default: 1.0)
"""
def __init__(self, params, lr=0.1, max_norm=0.01, stddev=2.0):
self.lr = lr
self.max_norm = max_norm
self.stddev = stddev
super().__init__(params, dict())
def step(self):
"""Performs a single optimization step.
The function expects the gradients to have been computed by BackPACK
and the parameters to have a ``batch_l2`` and ``grad_batch`` attribute.
"""
l2_norms_all_params_list = []
for group in self.param_groups:
for p in group["params"]:
l2_norms_all_params_list.append(p.batch_l2)
l2_norms_all_params = torch.stack(l2_norms_all_params_list)
total_norms = torch.sqrt(torch.sum(l2_norms_all_params, dim=0))
scaling_factors = torch.clamp_max(total_norms / self.max_norm, 1.0)
for group in self.param_groups:
for p in group["params"]:
clipped_grads = p.grad_batch * make_broadcastable(
scaling_factors, p.grad_batch
)
clipped_grad = torch.sum(clipped_grads, dim=0)
noise_magnitude = self.stddev * self.max_norm
noise = torch.randn_like(clipped_grad) * noise_magnitude
perturbed_update = clipped_grad + noise
p.data.add_(-self.lr * perturbed_update)
Running and plotting
We can now run our optimizer on MNIST.
optimizer = DP_SGD(model.parameters(), lr=0.1, max_norm=0.01, stddev=2.0)
losses = []
accuracies = []
for epoch in range(NUM_EPOCHS):
for batch_idx, (x, y) in enumerate(mnist_dataloader):
x, y = x.to(DEVICE), y.to(DEVICE)
optimizer.zero_grad()
outputs = model(x)
loss = loss_function(outputs, y)
with backpack(BatchGrad(), BatchL2Grad()):
loss.backward()
optimizer.step()
# Logging
losses.append(loss.detach().item())
accuracies.append(accuracy(outputs, y))
if (batch_idx % PRINT_EVERY) == 0:
print(
"Epoch %3.d/%d Iteration %3.d " % (epoch, NUM_EPOCHS, batch_idx)
+ "Minibatch Loss %.3f " % losses[-1]
+ "Accuracy %.3f" % accuracies[-1]
)
if MAX_ITER is not None and batch_idx > MAX_ITER:
break
Epoch 0/1 Iteration 0 Minibatch Loss 2.309 Accuracy 0.094
Epoch 0/1 Iteration 50 Minibatch Loss 1.543 Accuracy 0.562
Epoch 0/1 Iteration 100 Minibatch Loss 0.286 Accuracy 0.891
Epoch 0/1 Iteration 150 Minibatch Loss 0.259 Accuracy 0.906
Epoch 0/1 Iteration 200 Minibatch Loss 0.261 Accuracy 0.906
fig = plt.figure()
axes = [fig.add_subplot(1, 2, 1), fig.add_subplot(1, 2, 2)]
axes[0].plot(losses)
axes[0].set_title("Loss")
axes[0].set_xlabel("Iteration")
axes[1].plot(accuracies)
axes[1].set_title("Accuracy")
axes[1].set_xlabel("Iteration")
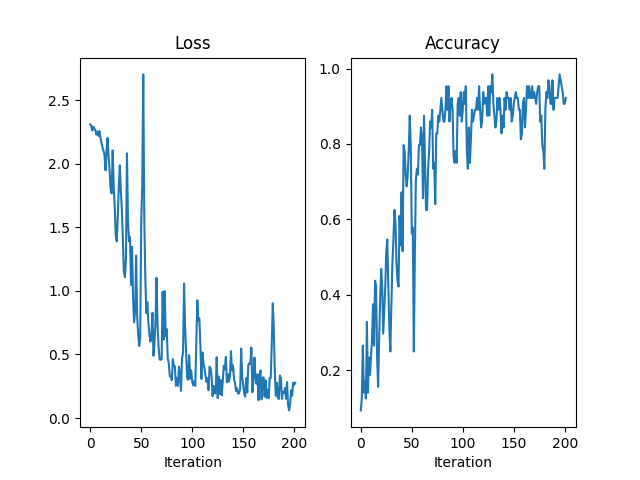
Text(0.5, 23.52222222222222, 'Iteration')
Total running time of the script: (0 minutes 10.953 seconds)