Note
Go to the end to download the full example code
Batched Jacobians
In PyTorch, you can easily compute derivatives of a scalar-valued variable
f w.r.t. to a variable param by calling
f.backward(). This computes the Jacobian ∂(f) / ∂(param)
that has shape [1, *param.shape].
If f is a reduction of a batched scalar fs of shape
[N], then BackPACK is capable to compute the individual gradients for
each scalar with its BatchGrad extension. This yields the Jacobian
∂(fs) / ∂(param) of shape [N, *param.shape].
This example demonstrates how to compute the Jacobian of a tensor-valued
variable fs, here for the example of a batched vector of shape
[N, C], whose Jacobian has shape [N, C, *param.shape].
Setup
We will use the batched vector-valued output of a simple MLP as tensor
fs that should be differentiated w.r.t. the model parameters
param_1, param_2, .... For param_i, this leads to a Jacobian
∂(fs) / ∂(param_i) of shape [N, C, *param_i.shape].
Let’s start by importing the required functionality and write a setup function to create our synthetic data.
import itertools
from math import sqrt
from typing import List, Tuple
import matplotlib.pyplot as plt
from torch import Tensor, allclose, cat, manual_seed, rand, zeros, zeros_like
from torch.autograd import grad
from torch.nn import Linear, MSELoss, ReLU, Sequential
from backpack import backpack, extend, extensions
# architecture specifications
N = 15
D_in = 10
D_hidden = 7
C = 5
def setup() -> Tuple[Sequential, Tensor]:
"""Create a simple MLP with ReLU activations and its synthetic input.
Returns:
A simple MLP and a tensor that can be fed to it.
"""
X = rand(N, D_in)
model = Sequential(Linear(D_in, D_hidden), ReLU(), Linear(D_hidden, C))
return model, X
With autograd
First, let’s compute the Jacobians with PyTorch’s autograd to verify
our results.
To do that, we need to differentiate per component of fs. This means
that we will differentiate multiple times through its graph, therefore we
need to set retain_graph=True.
manual_seed(0)
model, X = setup()
fs = model(X)
autograd_jacobians = [zeros(fs.shape + param.shape) for param in model.parameters()]
for n, c in itertools.product(range(N), range(C)):
grads_n_c = grad(fs[n, c], model.parameters(), retain_graph=True)
for param_idx, param_grad in enumerate(grads_n_c):
autograd_jacobians[param_idx][n, c, :] = param_grad
Let’s visualize the Jacobians by flattening the dimensions stemming from
fs and from param_i, and by concatenating them along the
parameter dimensions:
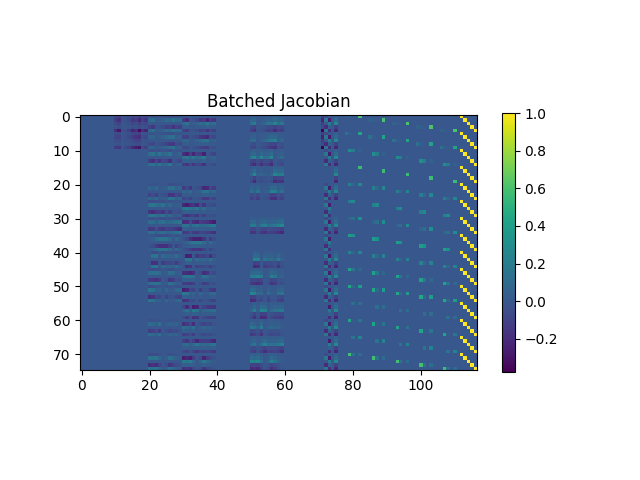
<matplotlib.colorbar.Colorbar object at 0x7efe5265f4f0>
In the following, we will compute the same Jacobian tensor lists with BackPACK. To compare our results, we will use the following helper function:
def compare_tensor_lists(
tensor_list1: List[Tensor], tensor_list_2: List[Tensor]
) -> None:
"""Checks equality of two tensor lists.
Args:
tensor_list1: First tensor list.
tensor_list2: Second tensor list.
Raises:
ValueError: If the two tensor lists don't match.
"""
if len(tensor_list1) != len(tensor_list_2):
raise ValueError("Tensor lists have different length.")
for tensor1, tensor2 in zip(tensor_list1, tensor_list_2):
if tensor1.shape != tensor2.shape:
raise ValueError("Tensors have different sizes.")
if not allclose(tensor1, tensor2):
raise ValueError("Tensors have different values.")
print("Both tensor lists match.")
Next, we will present two approaches to compute such Jacobians with BackPACK.
You can imagine the first one as carrying out the for-loop over N
parallel, and the second one as carrying out both for loops over N, C
in parallel. The first approach relies on a first-order extension, the second
one on a second-order extension. This means that while the first approach
works on quite general graphs, for the second one to work your graph must be
fully BackPACK-compatible.
With BackPACK’s BatchGrad
As described in the introduction, BackPACK’s BatchGrad extension can
compute Jacobians of batched scalars. We can therefore compute the
derivatives for fs[:, c] in one iteration, parallelizing the Jacobian
computation over the batch axis. For the full Jacobian, this requires
C backpropagations, hence we need to tell both autograd and
BackPACK to retain the graph.
Let’s do that in code, and check the result:
manual_seed(0)
model, X = setup()
model = extend(model)
fs = model(X)
backpack_first_jacobians = [zeros(fs.shape + p.shape) for p in model.parameters()]
for c in range(C):
with backpack(extensions.BatchGrad(), retain_graph=True):
f = fs[:, c].sum()
f.backward(retain_graph=True)
for param_idx, param in enumerate(model.parameters()):
backpack_first_jacobians[param_idx][:, c, :] = param.grad_batch
print("Comparing batched Jacobian from autograd with BackPACK (via BatchGrad):")
compare_tensor_lists(autograd_jacobians, backpack_first_jacobians)
Comparing batched Jacobian from autograd with BackPACK (via BatchGrad):
Both tensor lists match.
With BackPACK’s SqrtGGNExact
The second approach uses BackPACK’s SqrtGGNExact second-order
extension. It computes the matrix square root of the GGN/Fisher.
This approach uses that after feeding fs through a square loss with
reduction='sum', the GGN’s square root is the desired Jacobian up to
a normalization factor of √2 (to find out more, read Section 2 of [Dangel,
2021]), and a transposition due to
BackPACK’s internals.
Like that, we get the Jacobian in a single backward pass and don’t have to retain the graph:
manual_seed(0)
model, X = setup()
model = extend(model)
loss_func = extend(MSELoss(reduction="sum"))
fs = model(X)
fs_labels = zeros_like(fs) # can contain arbitrary values.
loss = loss_func(fs, fs_labels)
with backpack(extensions.SqrtGGNExact()):
loss.backward()
backpack_second_jacobians = [
param.sqrt_ggn_exact.transpose(0, 1) / sqrt(2) for param in model.parameters()
]
print("Comparing batched Jacobian from autograd with BackPACK (via SqrtGGNExact):")
compare_tensor_lists(autograd_jacobians, backpack_second_jacobians)
Comparing batched Jacobian from autograd with BackPACK (via SqrtGGNExact):
Both tensor lists match.
Total running time of the script: (0 minutes 0.226 seconds)