Note
Click here to download the full example code
Matrix-free second-order optimization¶
This example walks you through a second-order optimizer that uses the conjugate gradient (CG) method and matrix-free multiplication with the block diagonal of different curvature matrices to solve for the Newton step.
The optimizer is tested on the classic MNIST example from PyTorch. In particular, we will use a model that suffers from the vanishing gradient problem and is hence difficult to optimizer for gradient descent. Second-order methods are less affected by that issue and can train these models, as they rescale the gradient according to the local curvature.
A local quadratic model of the loss defined by a curvature matrix \(C(x_t)\) (the Hessian, generalized Gauss-Newton, or other approximations)is minimized by taking the Newton step
where
Let’s get the imports, configuration and some helper functions out of the way first. Notice that we are choosing a net with many sigmoids to make it hard to train for SGD.
Note
Larger batch sizes are usually recommended for second-order methods. However, the memory constraints imposed by the architecture used to build this example restrict us to rather small values.
import math
import matplotlib.pyplot as plt
import torch
from backpack import backpack, extend, extensions
from backpack.utils.examples import get_mnist_dataloder
BATCH_SIZE = 64
LR = 0.1
DAMPING = 1e-2
CG_TOL = 0.1
CG_ATOL = 1e-6
CG_MAX_ITER = 20
MAX_ITER = 50
PRINT_EVERY = 10
DEVICE = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
torch.manual_seed(0)
mnist_loader = get_mnist_dataloder(batch_size=BATCH_SIZE)
def make_model():
return torch.nn.Sequential(
torch.nn.Conv2d(1, 10, 5, 1),
torch.nn.Sigmoid(),
torch.nn.MaxPool2d(2, 2),
torch.nn.Conv2d(10, 20, 5, 1),
torch.nn.Sigmoid(),
torch.nn.MaxPool2d(2, 2),
torch.nn.Flatten(),
torch.nn.Linear(4 * 4 * 20, 50),
torch.nn.Sigmoid(),
torch.nn.Linear(50, 10),
)
model = make_model().to(DEVICE)
loss_function = torch.nn.CrossEntropyLoss().to(DEVICE)
def get_accuracy(output, targets):
"""Helper function to print the accuracy"""
predictions = output.argmax(dim=1, keepdim=True).view_as(targets)
return predictions.eq(targets).float().mean().item()
Writing the optimizer¶
To compute the update, we need access to the curvature matrix in form of matrix-vector products. We can then solve the linear system implied by the Newton step with CG,
and perform the update
for every parameter.
Here is the optimizer. At its core is a simple implementation of CG that
will iterate until the residual norm decreases a certain threshold (determined
the atol and tol arguments), or exceeds a maximum budget (maxiter).
class CGNOptimizer(torch.optim.Optimizer):
def __init__(
self,
parameters,
bp_extension,
lr=0.1,
damping=1e-2,
maxiter=100,
tol=1e-1,
atol=1e-8,
):
super().__init__(
parameters,
dict(
lr=lr,
damping=damping,
maxiter=maxiter,
tol=tol,
atol=atol,
savefield=bp_extension.savefield,
),
)
self.bp_extension = bp_extension
def step(self):
for group in self.param_groups:
for p in group["params"]:
damped_curvature = self.damped_matvec(
p, group["damping"], group["savefield"]
)
direction, info = self.cg(
damped_curvature,
-p.grad.data,
maxiter=group["maxiter"],
tol=group["tol"],
atol=group["atol"],
)
p.data.add_(direction, alpha=group["lr"])
def damped_matvec(self, param, damping, savefield):
curvprod_fn = getattr(param, savefield)
def matvec(v):
v = v.unsqueeze(0)
result = damping * v + curvprod_fn(v)
return result.squeeze(0)
return matvec
@staticmethod
def cg(A, b, x0=None, maxiter=None, tol=1e-5, atol=1e-8):
r"""Solve :math:`Ax = b` for :math:`x` using conjugate gradient.
The interface is similar to CG provided by :code:`scipy.sparse.linalg.cg`.
The main iteration loop follows the pseudo code from Wikipedia:
https://en.wikipedia.org/w/index.php?title=Conjugate_gradient_method&oldid=855450922
Parameters
----------
A : function
Function implementing matrix-vector multiplication by `A`.
b : torch.Tensor
Right-hand side of the linear system.
x0 : torch.Tensor
Initialization estimate.
atol: float
Absolute tolerance to accept convergence. Stop if
:math:`|| A x - b || <` `atol`
tol: float
Relative tolerance to accept convergence. Stop if
:math:`|| A x - b || / || b || <` `tol`.
maxiter: int
Maximum number of iterations.
Returns
-------
x (torch.Tensor): Approximate solution :math:`x` of the linear system
info (int): Provides convergence information, if CG converges info
corresponds to numiter, otherwise info is set to zero.
"""
maxiter = b.numel() if maxiter is None else min(maxiter, b.numel())
x = torch.zeros_like(b) if x0 is None else x0
# initialize parameters
r = (b - A(x)).detach()
p = r.clone()
rs_old = (r ** 2).sum().item()
# stopping criterion
norm_bound = max([tol * torch.norm(b).item(), atol])
def converged(rs, numiter):
"""Check whether CG stops (convergence or steps exceeded)."""
norm_converged = norm_bound > math.sqrt(rs)
info = numiter if norm_converged else 0
iters_exceeded = numiter > maxiter
return (norm_converged or iters_exceeded), info
# iterate
iterations = 0
while True:
Ap = A(p).detach()
alpha = rs_old / (p * Ap).sum().item()
x.add_(p, alpha=alpha)
r.sub_(Ap, alpha=alpha)
rs_new = (r ** 2).sum().item()
iterations += 1
stop, info = converged(rs_new, iterations)
if stop:
return x, info
p.mul_(rs_new / rs_old)
p.add_(r)
rs_old = rs_new
Running and plotting¶
Let’s try the Newton-style CG optimizer with the generalized Gauss-Newton (GGN) as curvature matrix.
After extend-ing the model and the loss function and creating the optimizer,
we have to add the curvature-matrix product extension to the with backpack(...)
context in a backward pass, such that the optimizer has access to the GGN product.
The rest is just a canonical training loop which logs and visualizes training
loss and accuracy.
model = extend(model)
loss_function = extend(loss_function)
optimizer = CGNOptimizer(
model.parameters(),
extensions.GGNMP(),
lr=LR,
damping=DAMPING,
maxiter=CG_MAX_ITER,
tol=CG_TOL,
atol=CG_ATOL,
)
losses = []
accuracies = []
for batch_idx, (x, y) in enumerate(mnist_loader):
optimizer.zero_grad()
x, y = x.to(DEVICE), y.to(DEVICE)
outputs = model(x)
loss = loss_function(outputs, y)
with backpack(optimizer.bp_extension):
loss.backward()
optimizer.step()
# Logging
losses.append(loss.detach().item())
accuracies.append(get_accuracy(outputs, y))
if (batch_idx % PRINT_EVERY) == 0:
print(
"Iteration %3.d/%3.d " % (batch_idx, MAX_ITER)
+ "Minibatch Loss %.5f " % losses[-1]
+ "Accuracy %.5f" % accuracies[-1]
)
if MAX_ITER is not None and batch_idx > MAX_ITER:
break
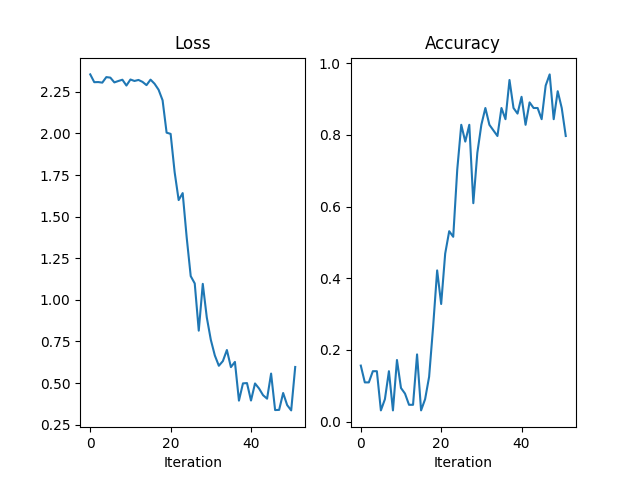
fig = plt.figure()
axes = [fig.add_subplot(1, 2, 1), fig.add_subplot(1, 2, 2)]
axes[0].plot(losses)
axes[0].set_title("Loss")
axes[0].set_xlabel("Iteration")
axes[1].plot(accuracies)
axes[1].set_title("Accuracy")
axes[1].set_xlabel("Iteration")
Out:
/home/docs/checkouts/readthedocs.org/user_builds/backpack/envs/1.2.0/lib/python3.7/site-packages/torch/nn/modules/module.py:795: UserWarning: Using a non-full backward hook when the forward contains multiple autograd Nodes is deprecated and will be removed in future versions. This hook will be missing some grad_input. Please use register_full_backward_hook to get the documented behavior.
warnings.warn("Using a non-full backward hook when the forward contains multiple autograd Nodes "
Iteration 0/ 50 Minibatch Loss 2.35480 Accuracy 0.15625
Iteration 10/ 50 Minibatch Loss 2.32407 Accuracy 0.09375
Iteration 20/ 50 Minibatch Loss 1.99725 Accuracy 0.32812
Iteration 30/ 50 Minibatch Loss 0.75986 Accuracy 0.82812
Iteration 40/ 50 Minibatch Loss 0.39462 Accuracy 0.90625
Iteration 50/ 50 Minibatch Loss 0.33532 Accuracy 0.87500
Text(0.5, 23.52222222222222, 'Iteration')
Vanishing gradients: comparison with SGD¶
By intention, we chose a model that is different to optimize with gradient descent due to the large number of sigmoids that reduce the gradient signal in backpropagation.
To verify that, let’s compare the Newton optimizer for different curvatures with SGD.
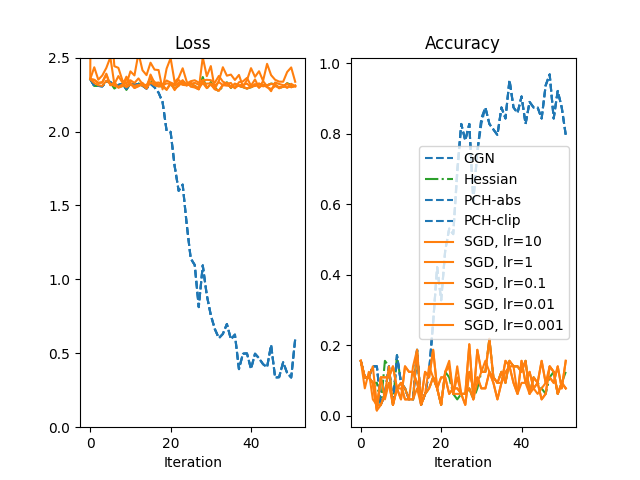
SGD is run for a large range of learning rates lr ∈ [10, 1, 0.1, 0.01, 0.001].
The performance of CG-Newton versus SGD is shown below (using a somewhat simplified color scheme to simplify the visualization).
def make_cgn_optimizer_fn(extension):
def optimizer_fn(model):
return CGNOptimizer(
model.parameters(),
extension,
lr=LR,
damping=DAMPING,
maxiter=CG_MAX_ITER,
tol=CG_TOL,
atol=CG_ATOL,
)
return optimizer_fn
curvatures = [
extensions.GGNMP(),
extensions.HMP(),
extensions.PCHMP(modify="abs"),
extensions.PCHMP(modify="clip"),
]
labels = [
"GGN",
"Hessian",
"PCH-abs",
"PCH-clip",
]
optimizers = []
for curvature in curvatures:
optimizers.append(make_cgn_optimizer_fn(curvature))
def make_sgd_optimizer_fn(lr):
def optimizer_fn(model):
return torch.optim.SGD(model.parameters(), lr=lr)
return optimizer_fn
sgd_lrs = [
10,
1,
0.1,
0.01,
0.001,
]
for lr in sgd_lrs:
optimizers.append(make_sgd_optimizer_fn(lr))
labels.append("SGD, lr={}".format(lr))
def train(optim_fn):
torch.manual_seed(0)
mnist_loader = get_mnist_dataloder(batch_size=BATCH_SIZE)
model = make_model().to(DEVICE)
loss_function = torch.nn.CrossEntropyLoss().to(DEVICE)
optimizer = optim_fn(model)
need_backpack = isinstance(optimizer, CGNOptimizer)
if need_backpack:
model = extend(model)
loss_function = extend(loss_function)
losses = []
accuracies = []
for batch_idx, (x, y) in enumerate(mnist_loader):
optimizer.zero_grad()
x, y = x.to(DEVICE), y.to(DEVICE)
outputs = model(x)
loss = loss_function(outputs, y)
if need_backpack:
with backpack(optimizer.bp_extension):
loss.backward()
else:
loss.backward()
optimizer.step()
# Logging
losses.append(loss.detach().item())
accuracies.append(get_accuracy(outputs, y))
if (batch_idx % PRINT_EVERY) == 0:
print(
"Iteration %3.d/%3.d " % (batch_idx, MAX_ITER)
+ "Minibatch Loss %.5f " % losses[-1]
+ "Accuracy %.5f" % accuracies[-1]
)
if MAX_ITER is not None and batch_idx > MAX_ITER:
break
return losses, accuracies
fig = plt.figure()
axes = [fig.add_subplot(1, 2, 1), fig.add_subplot(1, 2, 2)]
axes[0].set_title("Loss")
axes[0].set_ylim(0, 2.5)
axes[0].set_xlabel("Iteration")
axes[1].set_title("Accuracy")
axes[1].set_xlabel("Iteration")
for optim_fn, label in zip(optimizers, labels):
print(label)
losses, accuracies = train(optim_fn)
if "SGD" in label:
axes[0].plot(losses, "-", color="tab:orange", label=label)
axes[1].plot(accuracies, "-", color="tab:orange", label=label)
elif "Hessian" in label:
axes[0].plot(losses, "-.", color="tab:green", label=label)
axes[1].plot(accuracies, "-.", color="tab:green", label=label)
else:
axes[0].plot(losses, "--", color="tab:blue", label=label)
axes[1].plot(accuracies, "--", color="tab:blue", label=label)
plt.legend()
Out:
GGN
/home/docs/checkouts/readthedocs.org/user_builds/backpack/envs/1.2.0/lib/python3.7/site-packages/torch/nn/modules/module.py:795: UserWarning: Using a non-full backward hook when the forward contains multiple autograd Nodes is deprecated and will be removed in future versions. This hook will be missing some grad_input. Please use register_full_backward_hook to get the documented behavior.
warnings.warn("Using a non-full backward hook when the forward contains multiple autograd Nodes "
Iteration 0/ 50 Minibatch Loss 2.35480 Accuracy 0.15625
Iteration 10/ 50 Minibatch Loss 2.32407 Accuracy 0.09375
Iteration 20/ 50 Minibatch Loss 1.99725 Accuracy 0.32812
Iteration 30/ 50 Minibatch Loss 0.75986 Accuracy 0.82812
Iteration 40/ 50 Minibatch Loss 0.39462 Accuracy 0.90625
Iteration 50/ 50 Minibatch Loss 0.33532 Accuracy 0.87500
Hessian
Iteration 0/ 50 Minibatch Loss 2.35480 Accuracy 0.15625
Iteration 10/ 50 Minibatch Loss 2.32469 Accuracy 0.09375
Iteration 20/ 50 Minibatch Loss 2.32172 Accuracy 0.03125
Iteration 30/ 50 Minibatch Loss 2.33445 Accuracy 0.12500
Iteration 40/ 50 Minibatch Loss 2.29927 Accuracy 0.12500
Iteration 50/ 50 Minibatch Loss 2.31358 Accuracy 0.09375
PCH-abs
Iteration 0/ 50 Minibatch Loss 2.35480 Accuracy 0.15625
Iteration 10/ 50 Minibatch Loss 2.32406 Accuracy 0.09375
Iteration 20/ 50 Minibatch Loss 1.99709 Accuracy 0.32812
Iteration 30/ 50 Minibatch Loss 0.75543 Accuracy 0.84375
Iteration 40/ 50 Minibatch Loss 0.39352 Accuracy 0.90625
Iteration 50/ 50 Minibatch Loss 0.33675 Accuracy 0.87500
PCH-clip
Iteration 0/ 50 Minibatch Loss 2.35480 Accuracy 0.15625
Iteration 10/ 50 Minibatch Loss 2.32407 Accuracy 0.09375
Iteration 20/ 50 Minibatch Loss 1.99700 Accuracy 0.32812
Iteration 30/ 50 Minibatch Loss 0.75807 Accuracy 0.84375
Iteration 40/ 50 Minibatch Loss 0.39428 Accuracy 0.90625
Iteration 50/ 50 Minibatch Loss 0.33643 Accuracy 0.87500
SGD, lr=10
Iteration 0/ 50 Minibatch Loss 2.35480 Accuracy 0.15625
Iteration 10/ 50 Minibatch Loss 2.40687 Accuracy 0.07812
Iteration 20/ 50 Minibatch Loss 2.49646 Accuracy 0.03125
Iteration 30/ 50 Minibatch Loss 2.44262 Accuracy 0.12500
Iteration 40/ 50 Minibatch Loss 2.42848 Accuracy 0.09375
Iteration 50/ 50 Minibatch Loss 2.43353 Accuracy 0.09375
SGD, lr=1
Iteration 0/ 50 Minibatch Loss 2.35480 Accuracy 0.15625
Iteration 10/ 50 Minibatch Loss 2.34693 Accuracy 0.09375
Iteration 20/ 50 Minibatch Loss 2.32786 Accuracy 0.03125
Iteration 30/ 50 Minibatch Loss 2.31912 Accuracy 0.12500
Iteration 40/ 50 Minibatch Loss 2.30326 Accuracy 0.12500
Iteration 50/ 50 Minibatch Loss 2.29920 Accuracy 0.09375
SGD, lr=0.1
Iteration 0/ 50 Minibatch Loss 2.35480 Accuracy 0.15625
Iteration 10/ 50 Minibatch Loss 2.31614 Accuracy 0.09375
Iteration 20/ 50 Minibatch Loss 2.32291 Accuracy 0.03125
Iteration 30/ 50 Minibatch Loss 2.31921 Accuracy 0.12500
Iteration 40/ 50 Minibatch Loss 2.30273 Accuracy 0.12500
Iteration 50/ 50 Minibatch Loss 2.30004 Accuracy 0.09375
SGD, lr=0.01
Iteration 0/ 50 Minibatch Loss 2.35480 Accuracy 0.15625
Iteration 10/ 50 Minibatch Loss 2.32386 Accuracy 0.04688
Iteration 20/ 50 Minibatch Loss 2.32216 Accuracy 0.10938
Iteration 30/ 50 Minibatch Loss 2.32547 Accuracy 0.07812
Iteration 40/ 50 Minibatch Loss 2.29940 Accuracy 0.15625
Iteration 50/ 50 Minibatch Loss 2.30404 Accuracy 0.07812
SGD, lr=0.001
Iteration 0/ 50 Minibatch Loss 2.35480 Accuracy 0.15625
Iteration 10/ 50 Minibatch Loss 2.33067 Accuracy 0.04688
Iteration 20/ 50 Minibatch Loss 2.33119 Accuracy 0.10938
Iteration 30/ 50 Minibatch Loss 2.34716 Accuracy 0.07812
Iteration 40/ 50 Minibatch Loss 2.30991 Accuracy 0.15625
Iteration 50/ 50 Minibatch Loss 2.32288 Accuracy 0.07812
<matplotlib.legend.Legend object at 0x7fe8fb7d9c90>
While SGD is not capable to train this particular model, the second-order methods are still able to do so. Such methods may be interesting for optimization tasks that first-order methods struggle with.
Note that the Hessian of the net is not positive semi-definite. In this case, the local quadratic model does not have a global minimum and that complicates the usage of the Hessian in second-order optimization. This also provides a motivation for the other positive semi-definite Hessian approximations shown here.
Total running time of the script: ( 1 minutes 5.871 seconds)